Since 1987, the Association for Computing Machinery has awarded the annual Gordon Bell Prize to recognize outstanding achievements in high-performance computing (HPC). Presented each year at the International Conference for High-Performance Computing, Networking, Storage and Analysis (SC), the prizes not only reward innovative projects that employ HPC for applications in science, engineering, and large-scale data analytics but also provide a timeline of milestones in parallel computing.
As a frequent home to the world’s most powerful and smartest scientific supercomputers, the US Department of Energy’s (DOE’s) Oak Ridge National Laboratory (ORNL) has hosted many previous Gordon Bell honorees on its HPC systems. The Oak Ridge Leadership Computing Facility (OLCF), a DOE Office of Science User Facility located at ORNL, manages these systems and makes them available to scientists around the world to accelerate scientific discovery and engineering progress. Consequently, the OLCF has provided the HPC systems for 25 previous Gordon Bell Prize finalists and eight winners, including last year’s team from ETH Zürich.
This year, four projects that used ORNL’s IBM AC922 Summit supercomputer are finalists. The 2020 Gordon Bell Prize will be awarded November 19 at SC20. Here are the finalists that used Summit.
DeePMD-kit: A New Paradigm for Molecular Dynamics Modeling
“The code produced by Team DeePMD, with its ability to scale to huge numbers of atoms, while retaining chemical accuracy, is poised to transform the field of materials research. Applications to other fields will surely follow.”
—Michael Klein, Laura H. Camell Professor of Science, Temple University
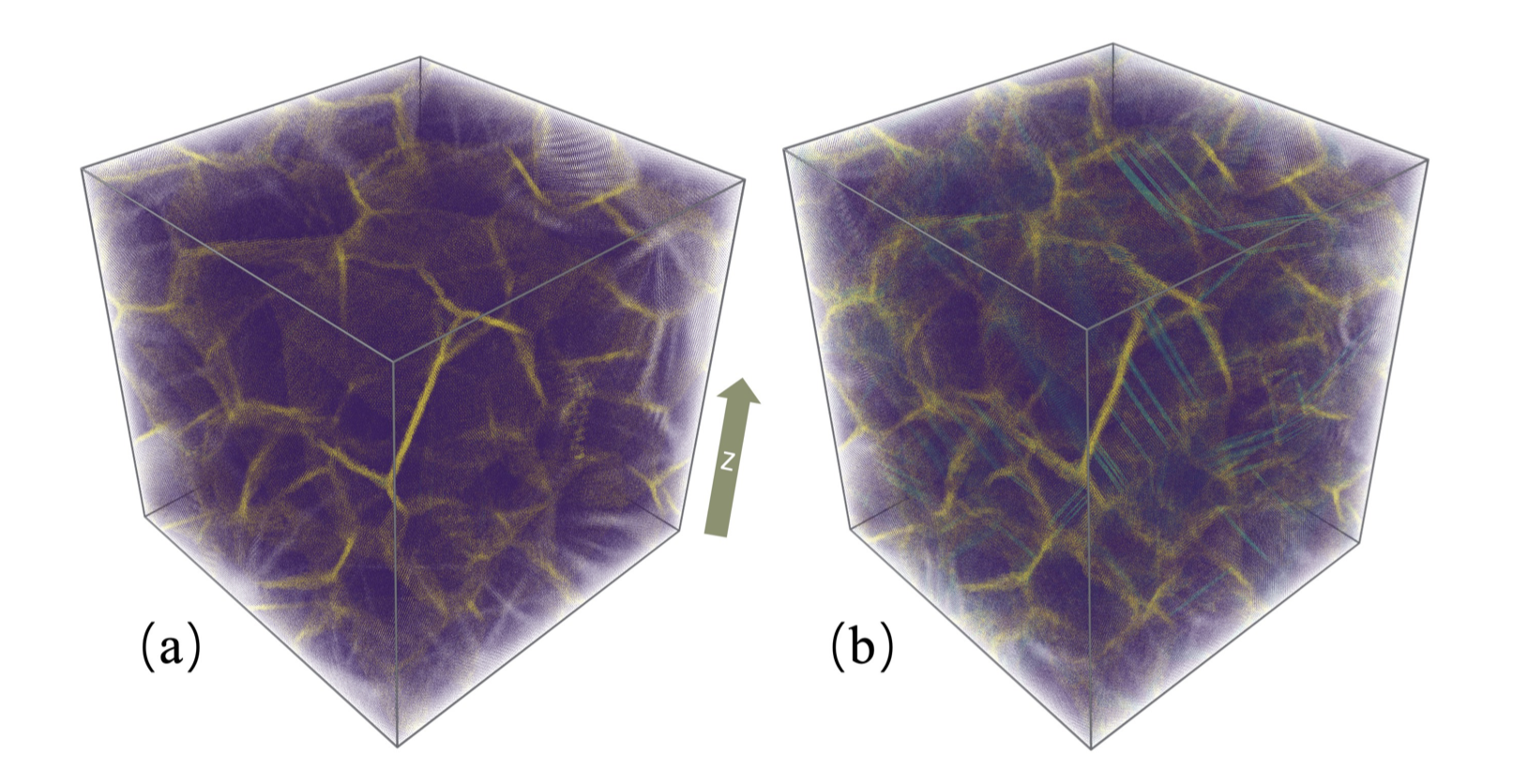
The DeePMD team simulated a block of copper with a system of 127.4 million atoms—more than 100 times larger current state of the art. Figure (a): A 10,401,218-atom nanocrystalline copper consisting of 64 randomly oriented crystals with 15-nm averaged grain diameter. Figure (b): The nanocrystalline copper after 10 percent tensile deformation along the z axis. Image Credit: DeePMD team
Molecular dynamics modeling has become a primary tool in scientific inquiry, allowing scientists to analyze the movements of interacting atoms over a set period of time, which helps them determine the properties of different materials or organisms. These computer simulations often lead the way in designing everything from new drugs to improved alloys. However, the two most popular methodologies come with caveats.
Classical molecular dynamics (MD), using Newtonian physics, can simulate trillions of particles on a modern supercomputer—however, its accuracy for more intricate simulations has limitations. Ab initio (“from the beginning”) molecular dynamics (AIMD), using quantum physics at each time step, can produce much more accurate results—but its inherent computational complexity limits the size and time span of its simulations. But what if there was a way to bridge the gap between MD and AIMD, to produce complex simulations that are both large and accurate?
With the power of ORNL’s Summit supercomputer, researchers from Lawrence Berkeley National Laboratory’s Computational Research Division; the University of California, Berkeley; the Institute of Applied Physics and Computational Mathematics, Peking University; and Princeton University successfully tested a software package that offers a potential solution: DeePMD-kit, named for “deep potential molecular dynamics.”
The team refers to DeePMD-kit as a “HPC+AI+Physical model” in that it combines high-performance computing (HPC), artificial intelligence (AI), and physical principles to achieve both speed and accuracy. It uses a neural network to assist its calculations by approximating the ab initio data, thereby reducing the computational complexity from cubic to linear scaling.
Simulating a block of copper atoms, the team put DeePMD-kit to the test on Summit with the goal of seeing how far they could push the simulation’s size and timescales beyond AIMD’s accepted limitations. They were able to simulate a system of 127.4 million atoms—more than 100 times larger than the current state of the art. Furthermore, the simulation achieved a time-to-solution mark of at least 1,000 times faster at 2.5 nanoseconds per day for mixed-half precision, with a peak performance of 275 petaflops (one thousand million million floating-point operations per second) for mixed-half precision.
“By combining physical principles and the representation power of deep neural networks, the Deep Potential method can achieve very good accuracy, especially for complex problems,” said Weile Jia, a postdoc in applied mathematics in Professor Lin Lin’s group at the Math Department of UC Berkeley, who co-led the project with Linfeng Zhang of Princeton. “Then we reorganize the data layout for bigger granularity on GPU and use data compression to significantly speedup the bottleneck. The neural network operators are optimized to the extreme, and most importantly, we successfully use half-precision in our code without losing accuracy.”
Related Publication: W. Jia, H. Wang, M. Chen, D. Lu, L. Lin, R. Car, W. E, and L. Zhang, “Pushing the limit of molecular dynamics with ab initio accuracy to 100 million atoms with machine learning,” International Conference for High Performance Computing, Networking, Storage and Analysis, SC20 (November 2020), arXiv:2005.00223
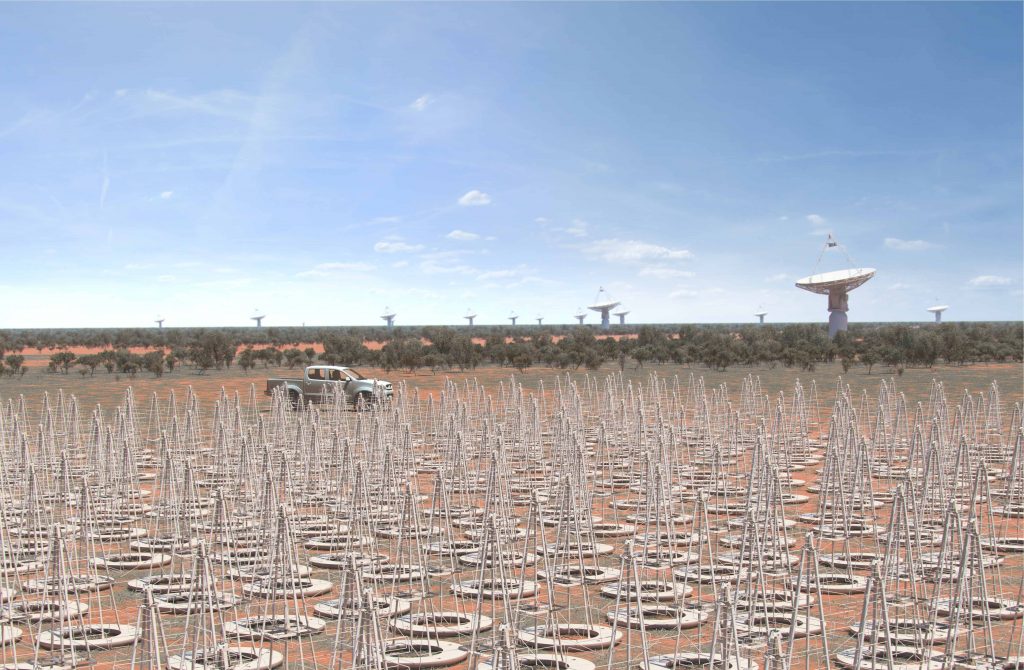

Once completed in the deserts of South Africa and Australia in the late 2020s, the Square Kilometre Array (SKA) will be a radio telescope array with a combined collecting area of over 1 square kilometer, or 1 million square meters. Image Credit: SKA
Square Kilometre Array: Massive Data Processing to Explore the Universe
“The innovative results already achieved and goals being pursued by this international team will greatly benefit the Next Generation Very Large Array, the Square Kilometre Array, and the next generation of radio interferometer facilities around the world.”
—Tony Beasley, Director, National Radio Astronomy Observatory
Scheduled to begin construction in 2021, the Square Kilometre Array (SKA) promises to become one of the biggest “Big Science” projects of all time (in physical size): a radio telescope array with a combined collecting area of over 1 square kilometer, or 1 million square meters. Once completed in the deserts of South Africa and Australia in the late 2020s, SKA’s thousands of dishes and low-frequency antennas will plumb the universe to figure out its mysteries.
SKA’s mission ultimately means it will produce massive amounts of information—an estimated 600 petabytes of data per year. Collecting, storing, and analyzing that data will be critical in producing SKA’s scientific discoveries. How will it be managed?
Building an end-to-end data-processing system on such an unprecedented scale is the task of an international team of radio astronomers, computer scientists, and software engineers. Workflow experts from the International Centre for Radio Astronomy Research (ICRAR) in Australia and the Shanghai Astronomical Observatory (SHAO) in China are developing the Daliuge workflow management system; GPU experts from Oxford University are optimizing the performance of the data generator; and input/output (I/O) experts at ORNL are producing I/O middleware based on the ORNL-developed Adaptable IO System (ADIOS). These three core software packages were completely developed by the team, with the original scope of running on top supercomputers.
Because SKA does not yet exist, its huge data output was simulated on Summit in order to test the team’s work, running a complete end-to-end workflow for a typical 6-hour SKA Phase 1 Low Frequency Array observation. The team used 99 percent of Summit, achieving 130 petaflops peak performance for single-precision, 247 gigabytes per second data generation rate, and 925 gigabytes per second pure I/O rate.
“For the first time, an end-to-end SKA data-processing workflow was executed in a production environment. It helps the SKA community—as well as the entire radio astronomy community—determine critical design factors for multi-billion-dollar next generation radio telescopes,” said Ruonan Wang, a software engineer in ORNL’s Scientific Data Group who works on the project. “It validated our ability, from both software and hardware perspectives, to process a key science case of SKA, which will answer some of the fundamental questions of our universe.”
Related Publication: R. Wang, A. Wicenec, and T. An, “Processing Full-Scale Square Kilometre Array Data on the Summit Supercomputer,” International Conference for High Performance Computing, Networking, Storage and Analysis, SC20 (November 202). doi: 10.1109/SC41405.2020.00006
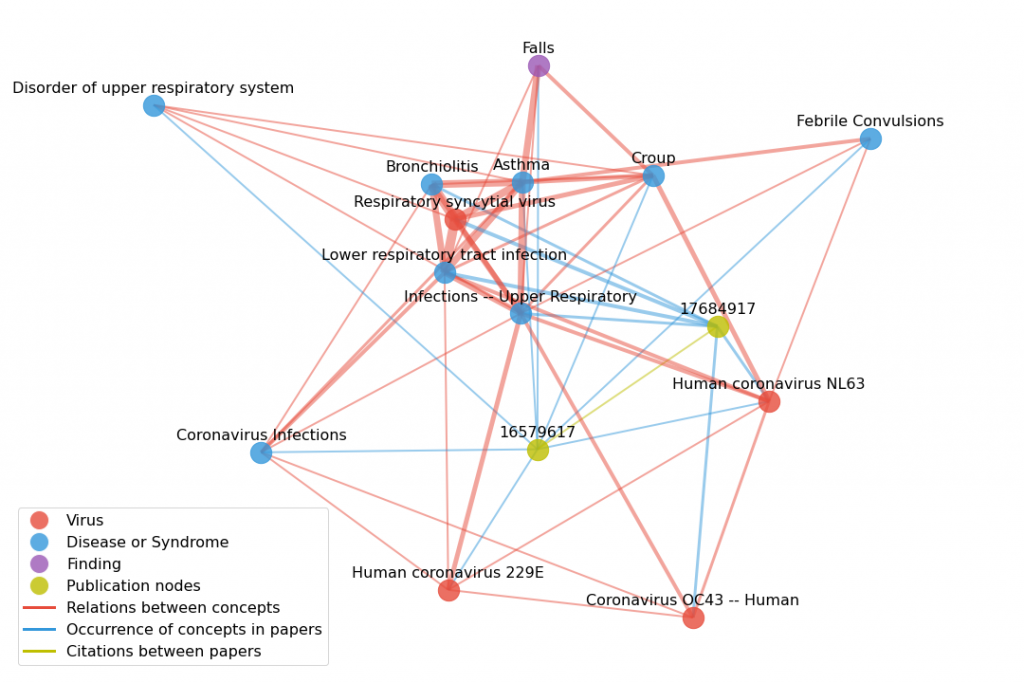
This is a small snapshot of the graph dataset used by DNSAPHOT team to conduct Swanson Linking for COVID-19 relations. The yellow nodes represent publications and the other nodes represents biomedical concepts such as viruses, diseases, or symptoms. The thickness of the edges and closeness of the nodes represents the likelihood of two concepts co-occurring. Image Credit: DSNAPSHOT team
DSNAPSHOT: An Accelerated Approach to Literature-Based Discovery
“The DSNAPSHOT algorithm approach … enables the identification of meaningful paths and novel relations on a previously unseen scale. Consequently, it moves the biomedical research community closer to a framework for analyzing how novel relations can be identified across the entire body of scientific literature.”
—Michael Weiner, PhD, VP AxioMx, Molecular Sciences and Head, Global Research of Abcam
In 1986, the late information scientist Don Swanson introduced the concept of “undiscovered public knowledge” in the field of biomedical research. His idea was both intriguing and straightforward: Out of the millions of published pieces of medical literature, what if there are yet unseen connections between their findings that could lead to new treatments? If, for example, “A affects B” in one study and “B affects C” in another, perhaps A and C have undiscovered commonalities worth investigating. Swanson proved his point by analyzing unrelated papers for such links, leading to hypothetical treatments that were later supported by clinical studies, such as taking magnesium supplements to help prevent migraine headaches. This process became known as “Swanson Linking.”
But in light of the enormous size of scientific literature in existence, mining it for undiscovered connections cannot be effectively conducted on a mass scale by mere humans. For example, the US National Library of Medicine’s PubMed database contains over 30 million citations and abstracts for biomedical literature. How can researchers possibly track that much information in its totality and find the patterns that may help identify new treatments?
One answer may be data-mining algorithms optimized for GPU-accelerated supercomputers such as ORNL’s Summit. When the federal government mobilized its national labs in the fight against COVID-19 in March, a team of ORNL and Georgia Tech researchers was assembled by ORNL computer scientist Ramakrishnan Kannan and Thomas E. Potok, head of ORNL’s Data and AI Systems Section of the Computer Science and Mathematics Division. The team’s mission was to investigate new ways of searching large-scale bodies of scholarly literature—and they ultimately found a way to conduct Swanson Linking on huge datasets at unprecedented speed.
Dasha Herrmannova from Kannan’s team began by creating a graph dataset based on Semantic MEDLINE—a dataset of biomedical concepts and the relations between them—extracted from PubMed. Then they expanded the graph with information extracted from the COVID-19 Open Research Dataset (CORD-19), resulting in a dataset of 18.5 million nodes representing concepts and papers, with 213 million relationships between them.
To search this massive dataset (via knowledge graph representations) for potential COVID-19 treatments, the team developed a new high-performance implementation of the Floyd-Warshall algorithm. The classic algorithm, originally published in 1962, determines the shortest distances between every pair of vertices in a given graph. (In terms of literature-based discovery, the shortest paths are usually more likely to reveal new connections between scholarly works.) Wanting to overcome the computational bottleneck of tackling massive graphs, Kannan, Piyush Sao, Hao Lu, and Robert Patton from ORNL, in collaboration with Vijay Thakkar and Rich Vuduc from Georgia Tech, optimized their version of the algorithm for distributed-memory parallel computers accelerated by GPUs. They named it Distributed Accelerated Semiring All-Pairs Shortest Path (DSNAPSHOT).
In effect, the team’s DSNAPSHOT is a supercharged version of Floyd-Warshall, able to identify the shortest paths in huge graphs in a matter of minutes. Using 90 percent of the Summit supercomputer—or 4,096 nodes, adding up to 24,576 GPUs—the team was able to compute an All-Pairs Shortest Path computation on a graph with 4.43 million vertices in 21.3 minutes. Peak performance reached 136 petaflops for single-precision. If every person on Earth completed one calculation per second, it would take the world’s population (~7 billion) 7 and a half months to complete what DSNAPSHOT can do in 1 second on Summit.
“To the best of our knowledge, DSNAPSHOT is the first method capable of calculating shortest path between all pairs of entities in a biomedical knowledge graph, thereby enabling the discovery of meaningful relations across the whole of biomedical knowledge,” Kannan said. “Looking forward, we believe this novel capability will enable the mining of scholarly knowledge corpora when embedded and integrated into artificial intelligence–driven natural language processing workflows at scale.”
Related Publication: R. Kannan, P. Sao, H. Lu, D. Herrmannova, V. Thakkar, R. Patton, R. Vuduc, and T. Potok, “Scalable Knowledge Graph Analytics at 136 Petaflop/s,” International Conference for High Performance Computing, Networking, Storage and Analysis, SC20 (November 2020).

Point defects in semiconductors are prototypes of solid-state qubits. Shown here is the isosurface enclosing 90 percent of the wave function for an in-gap state from a divacancy defect in silicon. Calculation cell contains 2,742 atoms and 10,968 electrons. Image Credit: BerkeleyGW team
BerkeleyGW: A New View into Excited-State Electrons
“The BerkeleyGW team’s demonstration of excited-state calculations with the GW method for 1,000-atom systems on accessible HPC facilities will be a game-changer. Researchers with diverse interests will be able to pursue fundamental understanding of excited states and physical processes in materials systems including novel two-dimensional semiconductors, electrochemical interfaces, organic molecular energy harvesting systems, and materials proposed for quantum information systems.”
—Mark S. Hybertsen, Group Leader, Theory & Computation Group Center for Functional Nanomaterials, Brookhaven National Laboratory
Historical epochs are often delineated by the materials that helped shape civilization, from the Stone Age to the Steel Age. Our current period is often referred to as the Silicon Age—but while those earlier eras were characterized by the structural properties of their predominant materials, silicon is different. Rather than ushering in new ways of building big things, its technological leap takes place on an atomic level, facilitating an information revolution.
Used as the main material in integrated circuits (AKA, the microchip), silicon has enabled the world of data processing we currently live in, from ever-more-powerful computers to unavoidable handheld devices. Central to its success has been the ability of chip designers to engineer these circuits to be increasingly faster and smaller, yet with more capacity as they add more and more transistors. But can microprocessor architects keep up with Moore’s law and continue to double the number of transistors in an integrated circuit every 2 years?
One route forward may be found in the work of a team of six physicists, materials scientists, and HPC specialists from the Berkeley Lab, UC Berkeley, and Stanford University that performed the largest-ever study of “excited-state” electrons using ORNL’s Summit supercomputer. Understanding and controlling such electronic excitation in silicon and other materials is key to designing the electronic and optoelectronic devices that have sparked the current information era. What’s more, the accurate modeling of excited-state properties of electrons in materials plays a crucial role in the rational design of other transformative technologies, including photovoltaics, batteries, and qubits for quantum information and quantum computing. In essence, the team’s high-performance calculations could help design new materials for these next generation technologies.
A state-of-the-art tool for determining excitations in materials is the “GW method,” an approach for calculating the self-energy (the quantum energy that a particle acquired from interactions with its surrounding environment) of a system of interacting electrons. The team adapted its own software package: BerkeleyGW—a quantum many-body perturbation theory code for excited states—to run on Summit’s GPU accelerators.
The team’s study of a system of defects in silicon and silicon carbide resulted in groundbreaking performance: the largest high-fidelity GW calculations ever made, with 10,986 valence electrons. By running on the entire Summit supercomputer, they also achieved 105.9 petaflops of double-precision performance with a time to solution of roughly 10 minutes.
“What’s really exciting about these numbers is that together they usher in the practical use of the high-fidelity GW method to the study of realistic complex materials,” said Jack Deslippe, team leader and head of the Applications Performance Group at the National Energy Research Scientific Computing Center, or NERSC. “These will be materials with defects, with interfaces, and with large geometries that drive real device design in quantum information, energy generation and storage, and next-gen electronics.”
Related Publication: M. Del Ben, C. Yang, Z. Li, F. H. da Jornada, S. G. Louie, J. Deslippe, “Accelerating Large-Scale Excited-State GW Calculations on Leadership HPC Systems,” International Conference for High Performance Computing, Networking, Storage and Analysis, SC20 (November 2020).
UT-Battelle LLC manages Oak Ridge National Laboratory for DOE’s Office of Science, the single largest supporter of basic research in the physical sciences in the United States. DOE’s Office of Science is working to address some of the most pressing challenges of our time. For more information, visit https://energy.gov/science.