Simulation provides a close-up look at the molecule that complicates next-generation biofuels
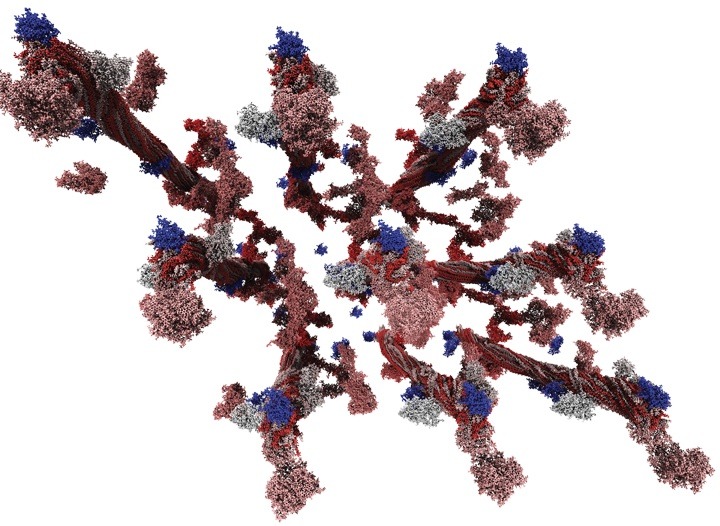
Leadership-class molecular dynamics simulation of the plant components lignin and cellulose. This 3.3 million-atom simulation was performed on 30,000 cores of the Jaguar XT5 supercomputer and investigated lignin precipitation on cellulose fibrils, a process that poses a significant obstacle to economically viable bioethanol production. Visualization by Mike Matheson, ORNL.
Lignin is very handy in many ways. In your diet it provides much of the fiber that keeps you happy and healthy. In a plant it helps keep the stalk and branches standing and strong. And in the future it may become the carbon fiber that makes cars and trucks lighter and more fuel efficient.
But for those who distill cellulosic ethanol, it’s just a bother.
A team led by Oak Ridge National Laboratory’s (ORNL’s) Jeremy Smith has taken a substantial step in the quest for cheaper biofuels by revealing the surface structure of lignin clumps down to 1 angstrom (equal to a 10 billionth of a meter, or smaller than the width of a carbon atom). The team’s conclusion, that the surface of these clumps is rough and folded, even magnified to the scale of individual molecules, was published June 15, 2011, in the journal Physical Review E.
Smith’s team employed two of ORNL’s signature strengths—simulation on Jaguar and neutron scattering—to resolve lignin’s structure at scales ranging from 1 to 1,000 angstroms.
Its results are important because lignin is a major impediment to the production of cellulosic ethanol, preventing enzymes from breaking down cellulose molecules into the sugars that will eventually be fermented.
Lignin itself is a very large, very complex molecule made up of hydrogen, oxygen, and carbon. In the wild its ability to protect cellulose from attack helps hardy plants such as switchgrass live in a wide range of environments. When these plants are used in biofuels, however, lignin is so effective that even expensive pretreatments fail to neutralize it.
The value of switchgrass
Switchgrass has many virtues as a source of ethanol, the primary renewable substitute for gasoline. It already grows wild throughout the country, it thrives in nearly any soil, and it appears to be happy in regions both wet and dry. Unlike traditional biofuel crops such as corn, switchgrass does not require constant care and attention, and it does not take up land and resources that would otherwise go toward producing food.
Yet the hardiness that allows switchgrass to thrive in inhospitable environments makes it stubbornly resistant to breakdown and fermentation.
“Nature has evolved a very sophisticated mechanism to protect plants against enzymatic attack,” explained team member Loukas Petridis, a computational physicist at ORNL, “so it is not easy to make the fuels. What we’re trying to do is understand the physical basis of biomass recalcitrance—resistance of the plants against enzymatic degradation.”
Switchgrass contains four major components: cellulose, lignin, hemicellulose, and pectin. The most important of these is cellulose, another large molecule, which is made up of hundreds to thousands of glucose sugar molecules strung together. In order for these sugars to be fermented, they must first be broken down in a process known as hydrolysis, in which enzymes move along and snip off the glucose molecules one by one.
Lignin blocks hydrolysis in two ways. First, it gets between the enzymes and the cellulose, forming a physical barrier. Second, it binds to passing enzymes, essentially taking them out of the game. Lignin does this job so well that ground-up switchgrass must be pretreated before it is hydrolyzed. After it is ground into small pieces, the switchgrass is heated above 300 degrees Fahrenheit in a dilute acid to make the cellulose more accessible to the enzymes.
Even then the lignin refuses to cooperate. While hemicellulose and pectin wash away with the pretreatment, the lignin re-forms into aggregates on the cellulose, droplets that can capture up to half the enzymes that are added to the mixture. These aggregates are the focus of Smith’s team’s study. The better that scientists are able to understand the aggregates, the better able they will be to design a more effective pretreatment process and find more successful enzymes.
According to Petridis, the team used neutron scattering with ORNL’s High Flux Isotope Reactor to resolve the lignin structure from 1,000 down to 10 angstroms. A molecular dynamics application called NAMD (for Not just Another Molecular Dynamics program) used Jaguar to resolve the structure from 100 angstroms down to 1. The overlap from 10 to 100 angstroms allowed the team to validate results between methods.
NAMD uses Newton’s law of motion to calculate the motion of a system of atoms—here lignin and water—typically in time steps of a femtosecond, or 1 thousand trillionth of a second. The two methods— neutrons and supercomputing—confirmed that the surface of lignin aggregates is highly folded, with a surface fractal dimension of about 2.6. The surface fractal dimension is a measure of the roughness or irregularity of a surface and ranges from 2 (very smooth) to 3 (very folded). The value of 2.6 is similar to that of broccoli. This roughness gives enzymes far more opportunity to get caught up in the lignin than a smooth surface would. In fact, a lignin droplet has about 31⁄2 times as much surface area as it would if it were smooth.
Smith’s project is the first to apply both molecular dynamics supercomputer simulations and neutron scattering to the structure of biomass. In fact, said Petridis, the two methods reinforce one another very well. On the one hand, neutron scattering could not reveal the structure at the smallest scales. On the other hand, simulation could not cover the full range of scales even on Jaguar, the United States’ most powerful supercomputer.
“When you look at the combination of neutrons and simulation, first of all it has not been done on lignins before. The combination of techniques gives you a multiscale picture of lignin. Neutron scattering can probe length scales, for example, from 10 angstroms all the way up to 1,000 angstroms.
“On the other hand, molecular dynamics simulations can go to smaller length scales—from 1 angstrom or even subangstrom all the way to 10 or even 100 angstroms. This is why we have been able to study the structure of the lignin droplets over various length scales. Not only was the finding new, but these techniques were used for the first time to study lignocellulose.”
While this research is an important step toward developing efficient, economically viable cellulosic ethanol production, much work remains. For example, this project focused only on lignin and included neither the cellulose nor the enzymes; in other words, it can tell us where the enzymes might fit on the lignin, but it has not yet told us whether the enzymes and lignin are likely to attract each other and attach.
Moving forward, the team is pursuing even larger simulations that include both lignin and cellulose. The latest simulations, on a 3.3 million-atom system, are being done with another molecular dynamics application called GROMACS (for Groningen Machine for Chemical Simulation).
This research and similar projects have the potential to make bioethanol production more efficient and less expensive in a variety of ways, Petridis noted. For example, earlier experiments showed that some enzymes are more likely to bind to lignin than others. The understanding of lignins provided by this latest research opens the door to further investigation into why that’s the case and how these differences can be exploited.
“To understand how this happens, you need an atomic-level or molecular-level description of both the enzymes, which we have already, and the lignin droplets, which we didn’t have until now. One thing we’d like to look at in the future is the interactions between the enzyme and the lignin. Of course, that would be a lot of work and require a lot of computer time.”
The research promises to be very enlightening, Petridis said, especially because it delves into areas that could not be fully explored before.
“Not knowing the structure of the droplets that play such an important role in biomass recalcitrance shows you that the field of biomass research has a lot of interesting questions to answer.”
Related publications:
J. Smith, et al. 2011. “Self-Similar Multiscale Structure of Lignin Revealed by Neutron Scattering and Molecular Dynamics Simulation,” Physical Review E, 83(6): 061911-1.