The “Pioneering Frontier” series features stories profiling the many talented ORNL employees behind the construction and operation of the OLCF’s incoming exascale supercomputer, Frontier. The HPE Cray system was delivered in 2021, with full user operations expected in 2022.
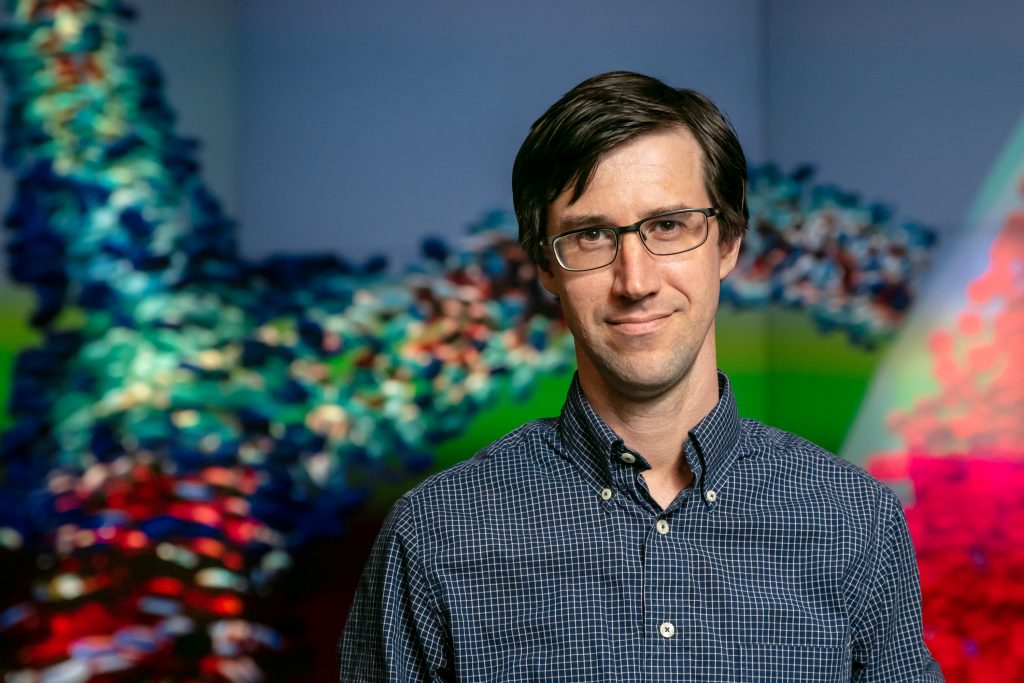

John Gounley is working on codes for the CANcer Distributed Learning Environment (CANDLE) project, which will provide deep learning methodologies to accelerate cancer research on machines like Frontier. Image Credit: Genevieve Martin, ORNL
Without scientific codes that scale, exascale would be little more than a number. That’s why computational scientists are working hard to ensure their codes run on exascale supercomputers as soon as these systems come online.
John Gounley is working on codes for the CANcer Distributed Learning Environment (CANDLE) project, which will provide deep learning methodologies to accelerate cancer research on machines like Frontier, the upcoming exascale system at the US Department of Energy’s (DOE’s) Oak Ridge National Laboratory (ORNL). Frontier will be capable of more than 1018 calculations when it is fully operational later this year. Formed out of a partnership between DOE and the National Cancer Institute (NCI), CANDLE is part of the Cancer Moonshot effort and exists within DOE’s Exascale Computing Project (ECP). Its objective is to develop applications from the pilot projects previously in the Cancer Moonshot effort, scale them up to next-generation supercomputers, and support their deep learning components.
“I work on the high-performance computing side of projects,” said Gounley, an ORNL computational scientist in the Advanced Computing for Health Sciences Section of the Computational Sciences and Engineering Division at ORNL. “I help take things we’ve done at small scales and get them up and running on big, leadership computing systems like Frontier.”
Gounley and his team have successfully run a “Transformer”—a deep learning model that identify unseen connections between words in clinical text—on Crusher, which is the testbed system for Frontier that contains identical hardware to that of the future exaflop system. Crusher is being used by the Center of Accelerated Application Readiness teams, ECP teams, staff at the National Center for Computational Sciences (NCCS), and the Oak Ridge Leadership Computing Facility’s (OLCF’s) vendor partners. The OLCF is a DOE Office of Science user facility located at ORNL.
“We’ve got our core software stack running on Crusher right now,” Gounley said. “It’s running a deep learning model called BERT [Bidirectional Encoder Representations from Transformers], which is a natural language processing [NLP] model.”
NLP models can process text in a similar way as humans. Gounley said BERT is both effective and accurate in the NLP space. However, it’s also computationally expensive to train.

Gounley works with CANDLE and another project called Modeling Outcomes using Surveillance data and Scalable AI for Cancer (MOSSAIC), which was originally one of the three pilot projects in the Cancer Moonshot effort. Image Credit: Genevieve Martin, ORNL
“We really need large high-performance computing resources to be able to run this effectively,” Gounley said.
Despite his current role as the technical lead for CANDLE, Gounley didn’t start out with an eye toward health research or supercomputing. Instead, he’s always been a numbers person.
“I wrote my first computer program to crunch numbers on high school sports statistics,” Gounley said.
When he was attending the Computational and Applied Mathematics graduate program at Old Dominion University, his work pivoted to health when he began modeling shape changes of red blood cells.
“I ended up doing a postdoctoral fellowship at Duke University with Amanda Randles, and she wanted to take the blood cells I was modeling, increase their computational size, and simulate hundreds of millions of cells,” Gounley said. “Naturally, that brings you into high-performance computing.”
Now, Gounley works with CANDLE and another project called Modeling Outcomes using Surveillance data and Scalable AI for Cancer (MOSSAIC), which was originally one of the three pilot projects in the Cancer Moonshot effort. NCCS Director Gina Tourassi is the principal investigator of MOSSAIC, and Heidi Hanson, group leader of the Biostatistics and Multiscale Systems Group in the Computational Sciences and Engineering Directorate at ORNL, is the technical lead.
For both the CANDLE and MOSSAIC efforts, Gounley’s team is working with deep learning models that are trained to pull out useful information from clinical text. When Gounley’s team trains a model on a large data set, it looks for patterns in the text, recognizes them when it sees them again, and appropriately classifies them. The team has now demonstrated that their models can accomplish this with a high degree of accuracy, and they ultimately aim to build even better deep learning models.
Frontier will enable the team to use a much larger neural language processing model with many more parameters, and they expect that it will be even more accurate.
“Having more parameters means we can recognize more patterns—or more complicated patterns—and that’s what we are expecting to be effective in the long term,” Gounley said.
Ultimately, Gounley’s team aims to provide NCI with better and more accurate cancer surveillance models.
“We hope that our next generation of models trained on systems like Frontier is going to be based on this BERT architecture and is going to be significantly more accurate than the models we have in practice today,” Gounley said.
Gounley said he is most excited about Frontier’s GPU memory capacity.
“Not only are the GPUs faster, but they’re much larger, in terms of the amount of memory they have,” Gounley said.
Frontier will have 128 gigabytes of memory per GPU, a fourfold increase from the NVIDIA Tesla V100 GPUs in the OLCF’s 200-petaflop IBM AC922 Summit supercomputer.
“That’s significant for us, because we make tradeoffs, with respect to model size, in that we really want the full model to fit on one GPU,” Gounley said. “You lose a lot of performance when your model goes off the GPU. So, if you’ve got a much larger GPU, a lot more things are possible.”
UT-Battelle LLC manages Oak Ridge National Laboratory for DOE’s Office of Science, the single largest supporter of basic research in the physical sciences in the United States. DOE’s Office of Science is working to address some of the most pressing challenges of our time. For more information, visit https://energy.gov/science.