A new paper published in Nature Communications adds further evidence to the bradykinin storm theory of COVID-19’s viral pathogenesis—a theory that was posited 2 years ago by a team of researchers led by Systems Biologist Dan Jacobson at the US Department of Energy’s (DOE’s) Oak Ridge National Laboratory (ORNL).
At the height of the pandemic, Jacobson and his team used ORNL’s Summit supercomputer to analyze gene-expression data of lung cells from COVID-19 patients. Their research suggested that genes related to some of the body’s systems that are responsible for controlling blood pressure, fluid balance, and inflammation—the renin angiotensin and kallikrein-kinin systems—appear to be excessively dysregulated (or impaired) in the lung cells of those infected with the virus. In a paper published in eLife, the team predicted that overproduction of bradykinin—the compound that dilates blood vessels and makes them permeable—could be the source of COVID-19 symptoms, such as excessive accumulation of fluid in the lungs, fatigue, nausea, and decreased cognitive function.
That theory has been further supported in a new study conducted by Jacobson and his colleagues in ORNL’s Biosciences, Computational Sciences and Engineering, and Neutron Scattering Divisions in collaboration with Soichi Wakatsuki, a professor of photon science at Stanford University’s SLAC National Accelerator Laboratory. Wakatsuki’s team was able to prove experimentally that the virus’s main protease, 3CLpro, binds to the NF-κB Essential Modulator (NEMO). The subsequent cleavage of NEMO means it dysregulates NF-κB, which is a protein complex that helps regulate the immune system’s response to infection—and its dysregulation can contribute to a bradykinin storm, just as the ORNL team’s pathogenesis model had predicted.
“This is the culmination of a lot of work coming from a lot of different angles,” Jacobson said. “We’re a computational systems biology group, so our previous work was really based on large-scale data analysis. This takes all of that computational work into the wet lab to generate new datasets to confirm the enzymatic activity and structural interactions. It’s incredibly exciting to see all these lines of evidence come together and then be validated—that everything our previous work was predicting to be the case is in fact true.”
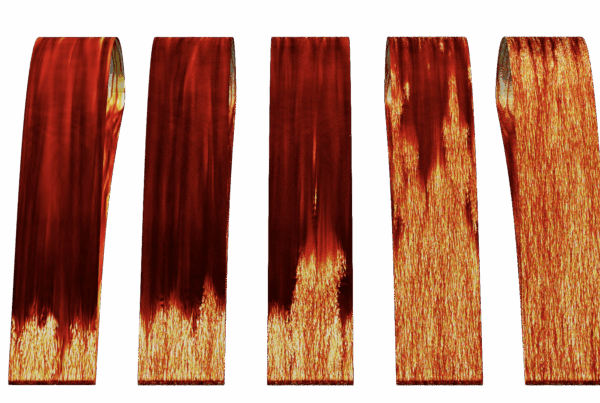
At SLAC, Wakatsuki’s team was able to use viral 3CLpro proteins (produced by ORNL senior scientist Andrey Kovalevsky and his team) and peptides to represent the cleavage sites in NEMO. The team then used X-ray crystallography to show the structural interaction between the two. Furthermore, a team at ORNL led by former ORNL researcher Stephanie Galanie was able to show, biochemically, that 3CLpro can cleave NEMO at physiologically relevant concentrations.
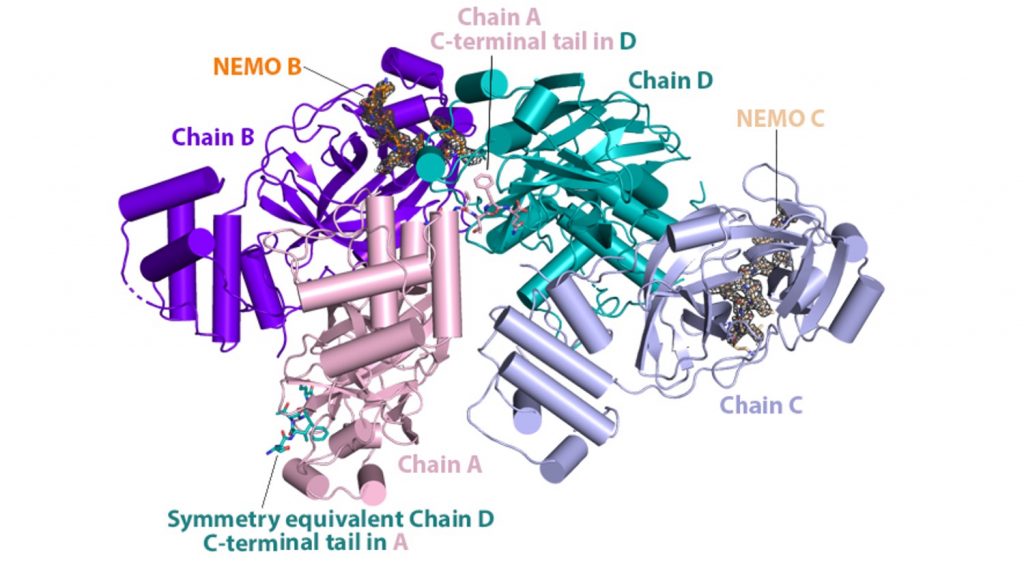
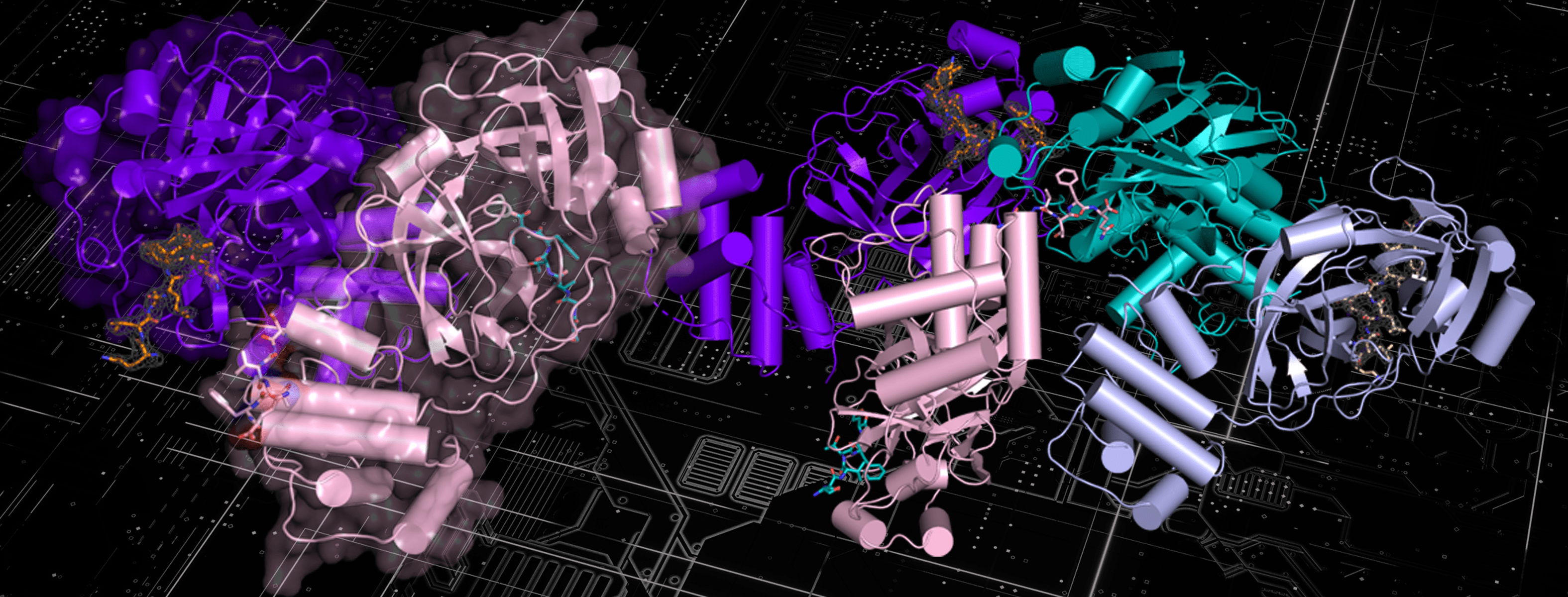
Shown here is the structure of the NEMO protein. A team from ORNL conducted extensive molecular dynamics work on Summit by using both quantum mechanics and machine-learning methods to look at the binding affinity of NEMO and 3CLpro in humans and other species and to consider the structural models derived from the sequences of other coronaviruses. Image courtesy Nature Communications, Dan Jacobson/ORNL.
“We now have atomistic-level evidence and biochemistry confirming the hypothesis that it binds and cleaves just how we expected it to,” Jacobson said.
This cross-lab collaboration at ORNL and SLAC came about through the National Virtual Biotechnology Laboratory,or NVBL, a DOE program funded by the Coronavirus Aid, Relief and Economic Security Act in 2020, that encouraged national labs in the fight against COVID-19. Wakatsuki and Jacobson met after Jacobson made a pitch at one of the NVBL virtual sessions and asked for collaborators to help prove his bradykinin storm theory through structural biology experiments.
“We went looking for people to do this next step with us, and Soichi spoke up at one of the meetings and said, ‘Yes, let’s go.’ And here we are now with a nice high-impact paper. I think that’s a real benefit of the collaborative approach that the NVBL had the national labs work together on, and I would like to see more of it,” Jacobson said.
As part of this effort, ORNL Computational Systems Biologist Erica Prates, then a postdoctoral researcher and now an early career staff member in the Biosciences Division, coordinated a team that included Omar Demerdash, Julie Mitchell, and Stephan Irle of ORNL. They conducted extensive molecular dynamics work on Summit by using both quantum mechanics and machine-learning methods to look at the binding affinity of NEMO and 3CLpro in humans and other species and to consider the structural models derived from the sequences of other coronaviruses.
“Erica is playing an important role in what we are calling structural systems biology to bridge across the computational efforts in the fields of systems biology and structural biology,” Jacobson said.
This team’s research will lead to a better understanding of the effects of different viruses, including zoonotic diseases (i.e., human diseases that originate from animals) in different host species. This knowledge will be vital in the effort to predict or even prevent the next pandemic.
“Our COVID work continues, but a big part of our focus has shifted toward pandemic prevention,” Jacobson said. “We have new funding [that we] obtained in collaboration with a number of other institutions for research that is really focused on dynamic prevention and trying to understand the rules of zoonosis and the effects, for example, of climate changes and how they are driving new zoonotic spillover events.”
Jacobson and his colleagues are currently partnering with Johns Hopkins University, Cornell University, and others to conduct a wide range of field studies and assays to analyze the interactions between viral proteins and host proteins, thereby creating the datasets needed for the computational models that will make virus predictions across whole ranges of species.
“Why do viruses happily live in some species nonpathogenically but become pathogens when zoonosis spillover occurs? How do they hop between different host species and be nonpathogenic until they hit humans?” Jacobson said. “The rules behind zoonosis are very poorly understood, and we have some really exciting work underway in which we’re building predictive models to understand the variables in the environment that can lead to these spillover events.”
The teams’ research was also partially funded from ORNL’s Laboratory Directed Research and Development Program, which supported the conceptual work on the NEMO cleavage in animal models for COVID-19 pathology. This work used DOE Office of Science user facilities including the Oak Ridge Leadership Computing Facility, the Spallation Neutron Source, and the High Flux Isotope Reactor, all at ORNL, and the Stanford Synchrotron Radiation Lightsource at SLAC.
Funding for human pathogenesis conceptualization was provided by a grant from the National Institutes of Health.
Learn more about ORNL research in the fight against COVID-19.
Related Publication: Mikhail Ali Hameedi, Erica T. Prates, Michael R. Garvin, Irimpan I. Mathews, B Kirtley Amos, Omar Demerdash, Mark Bechthold, Mamta Iyer, Simin Rahighi, Daniel W. Kneller, Andrey Kovalevsky, Stephan Irle, Van-Quan Vuong, Julie C Mitchell, Audrey Labbe, Stephanie Galanie, Soichi Wakatsuki, and Daniel Jacobson, “Structural and functional characterization of NEMO cleavage by SARS-CoV-2 3CLpro,” Nature Communications 13, (2022), https://doi.org/10.1038/s41467-022-32922-9.
UT-Battelle manages ORNL for DOE’s Office of Science, the single largest supporter of basic research in the physical sciences in the United States. DOE’s Office of Science is working to address some of the most pressing challenges of our time. For more information, visit https://energy.gov/science.