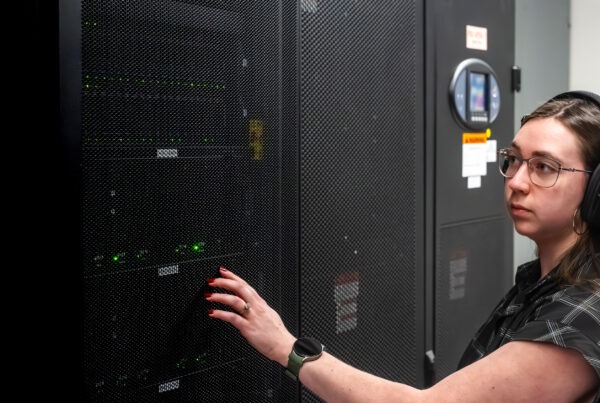
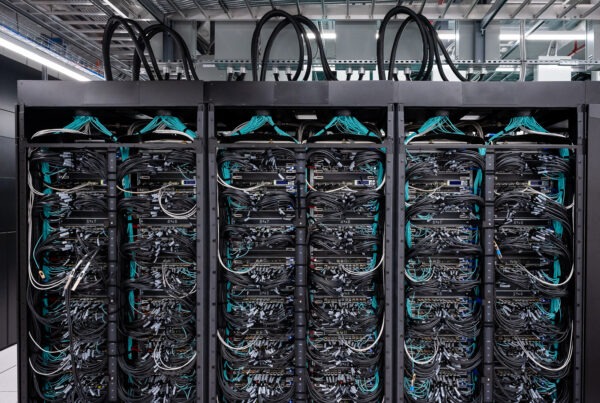
To those outside the high-performance computing community, supercomputers are a mysterious breed. Rows of refrigerator-sized cabinets span the length of an entire room, connected in an ensemble of fiber-optic cables and networking equipment. But one question remains: where is the monitor? As it turns out, the answer has a lot more to do with visualization tools than hardware.
Although supercomputers aren’t connected to a single monitor, visualization specialists like computer scientist Benjamín Hernández in the Advanced Data and Workflow Group at the Oak Ridge Leadership Computing Facility (OLCF), can take data produced by these large systems and visualize it scientifically using specialized software. And now, they may be able to do so with more ease than ever before.
A team at the OLCF recently joined an effort aimed at creating a standardized application programming interface (API) for a visualization method called ray tracing, which models the way light behaves to create highly physically accurate images from datasets. The move would allow new visualization software programs and custom visualization codes to more easily run on any computing device—even those with drastically different architecture
Creating physically accurate images allows users to intuitively understand complex datasets. The traditional technique for creating images—called rasterization—requires the implementation of multiple algorithms to create shadows, reflections, or other features. With ray tracing, these features are straightforward and trivial to generate. Combined with the recent ability to accelerate the technique using many-core and multicore processors, ray tracing has become an increasingly popular technique. Hernández performs it on the world’s most powerful and smartest supercomputer: the IBM AC922 Summit supercomputer at the OLCF, a US Department of Energy (DOE) Office of Science User Facility located at DOE’s Oak Ridge National Laboratory (ORNL). Using Summit’s GPUs, SIGHT—a visualization tool Hernández developed—is capable of rendering images through either rasterization or ray tracing, producing and streaming high-end visualizations that are sent back to the user’s workstation or laptop.

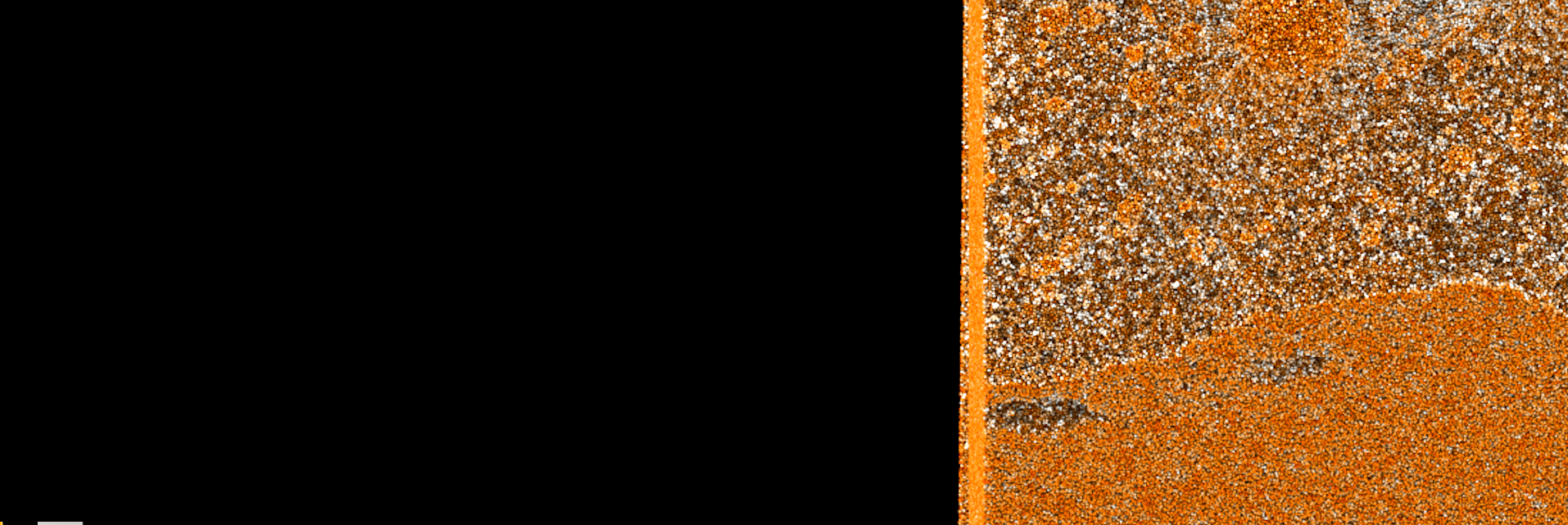
SIGHT’s ray-traced visualizations demonstrate the benefit of using shadows to enhance visual perception. SIGHT uses NVIDIA Optix and Intel OSPRay and is supported in the OLCF’s Summit and Rhea systems. Image Credit: Benjamín Hernández, ORNL
Ray tracing boasts many benefits and companies such as NVIDIA and Intel have made available architecture-specific libraries to render ray-traced images. However, this means developers using multiple supercomputers must determine how to best port their codes and support the technique on different architectures. For teams working on complex visualization codes such as Visual Molecular Dynamics (VMD), Paraview, or VisIt, the task becomes challenging.
“Historically, visualization teams have had to develop or deploy codes that talk to a proprietary API, deploy it on the specific compute cluster that uses that API, and then repeat the process for other systems and APIs,” Hernández said. “This effort focuses on enabling portability across different architectures, which is one of DOE’s main objectives when supporting these initiatives.”
Hernández is leading OLCF’s involvement with a new Analytic Rendering Exploratory Group within the Khronos Group that discusses the standardization of a cross-platform API for data visualization. The Khronos Group is an industry consortium focused on developing open interoperability standards for graphics rendering and parallel computation. Such an open, cross-platform analytic rendering API standard would give scientists the ability to easily reuse their visualization codes across diverse systems and processors.
“A unified API would abstract all the specific features related to different libraries and allow them to be used more widely,” said Mark Kim, a research scientist in the Scientific Data Group at ORNL who works with Hernández on the project. “It would allow developers to spend less time coding for different libraries and more time adding features that can benefit users and, ultimately, their science.”
If the effort is successful, the team believes the first version of the standard will be released in 2020.
“This emerging standard will greatly simplify the development of scientific visualization applications and enable a wider group of scientists to benefit from the latest advances in rendering technology,” said Peter Messmer, senior manager of High-Performance Computing Visualization at NVIDIA.
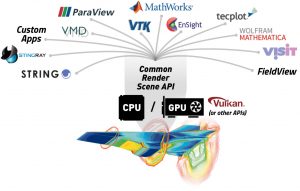
An analytic rendering API standard would lead to advances in code portability and reusability. Image Credit: Khronos Group
The benefits, Hernández said, will be multifaceted. Along with better renderings and more realistic images, researchers will be able to more easily produce images for journal covers and other communication purposes.
“We have three kinds of use cases for this: the researcher who needs to visualize scientific data, the developer of the visualization technology, and the deployment team for that visualization technology,” Hernández said.
Once implemented, Hernández said, the standard will benefit all three.
“A standard interface for using vendor-implemented rendering systems will reduce the amount of engineering effort spent on connecting state-of-the-art rendering systems to end-user applications and increase the available effort vendors can devote to increasing the value of their rendering systems for everyone,” said Jefferson Amstutz, lead OSPRay engineer at Intel.
Neil Trevett, Khronos Group president, believes the participation from code developers in concert with users will promote major progress in code portability.
“Khronos warmly welcomes ORNL’s participation in its Analytic Rendering Exploratory Group,” Trevett said. “The combination of developers and users of visualization technology working together is a powerful combination that will lead to genuine advances in code reusability and portability for the scientific community.”
UT Battelle LLC manages ORNL for DOE’s Office of Science. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States and is working to address some of the most pressing challenges of our time. For more information, please visit https://energy.gov/science.






